Accuracy
de novo assembly
dDocent employs a series of data reduction techniques, aligment based clustering (using CD-hit), and, for PE assembly, a specialized RAD assembly software (rainbow. This combination allows for accurate and effecient de novo assembly.
A comparison among pipelines
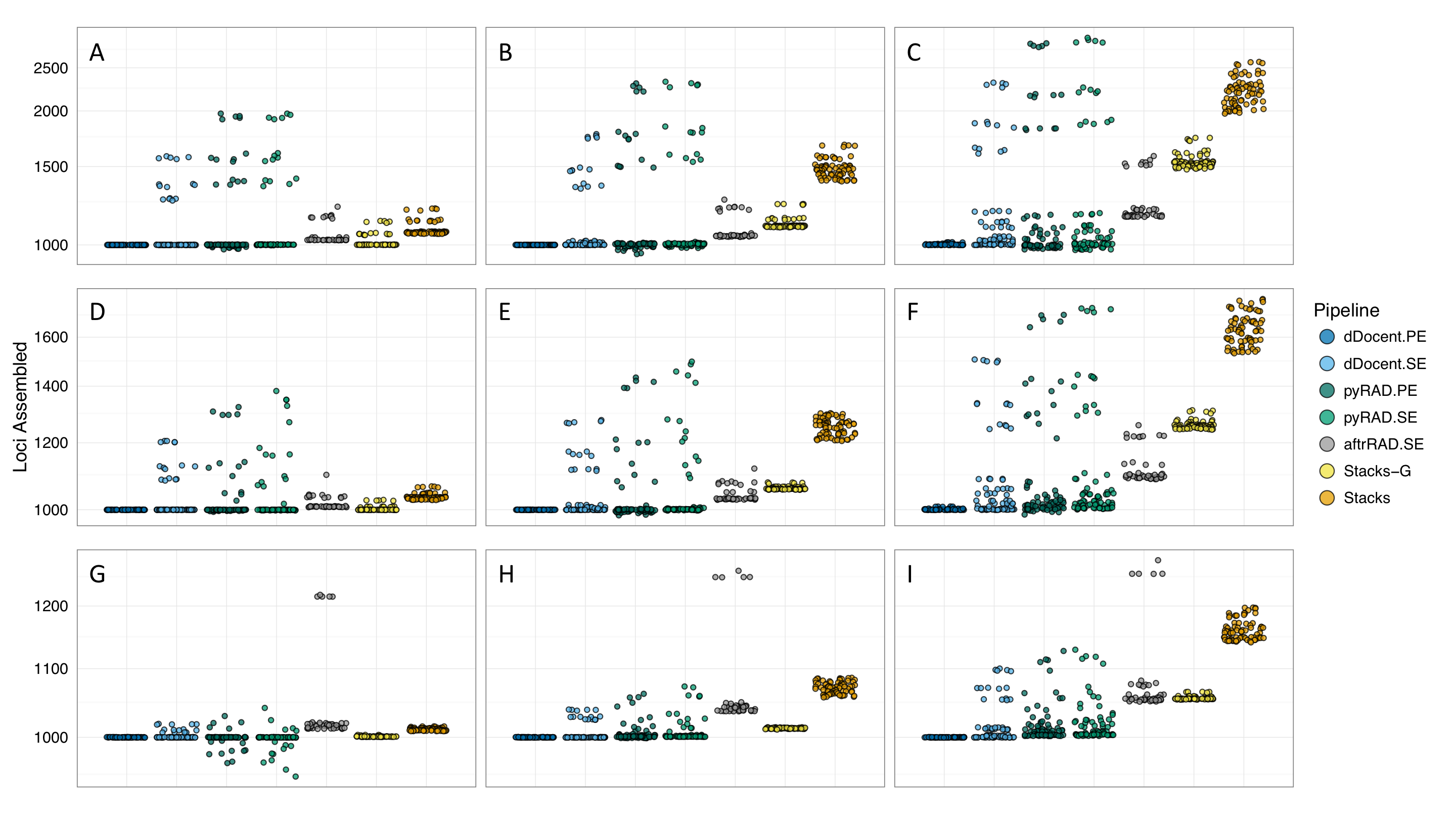
This is 1000 simulated ddRAD data loci being assembled across a variety of parameters for each pipeline.
Overclustering leads to bias
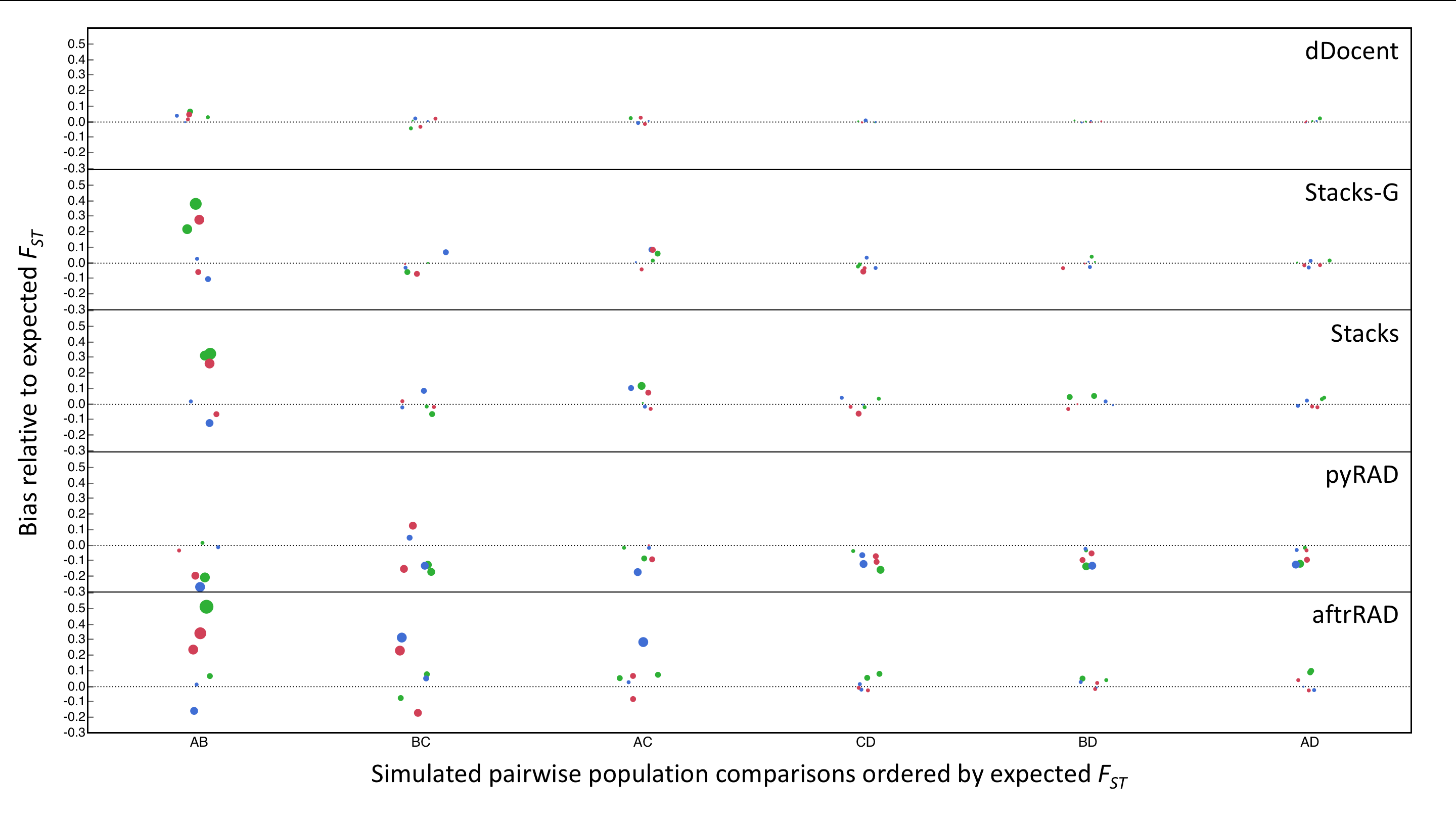
Above is a figure depicting the relative bias of pairwise FST values generated by different RADseq bioinformatic pipelines. Circles are sized according to the magnitude of bias (Observed - Expected)/Expected and are colored relative to the percentage of INDEL variation simulated in the data set: blue- 1%, red-5%, and green-10%. Simulations consisted of four populations in a stepping stone model with a decreasing migration rates.
Bayesian, haplotype based, population-aware, genotyping from FreeBayes
FreeBayes is a Bayesian genetic variant detector designed to detect SNPs, INDels (insertions and deletions), and complex events (composite insertion and substitution events) smaller than the length of a short-read sequencing alignment. FreeBayes is haplotype-based, in the sense that it calls variants based on the literal sequences of reads aligned to a particular target, not their precise alignment, and for any number of individuals from a population and a to determine the most-likely combination of genotypes for the population at each position in the reference.